Section: New Results
Data-driven HRF estimation for encoding and decoding models
Participants : Fabian Pedregosa Izquierdo [correspondant] , Michael Eickenberg, Alexandre Gramfort, Philippe Ciuciu, Bertrand Thirion, Gaël Varoquaux.
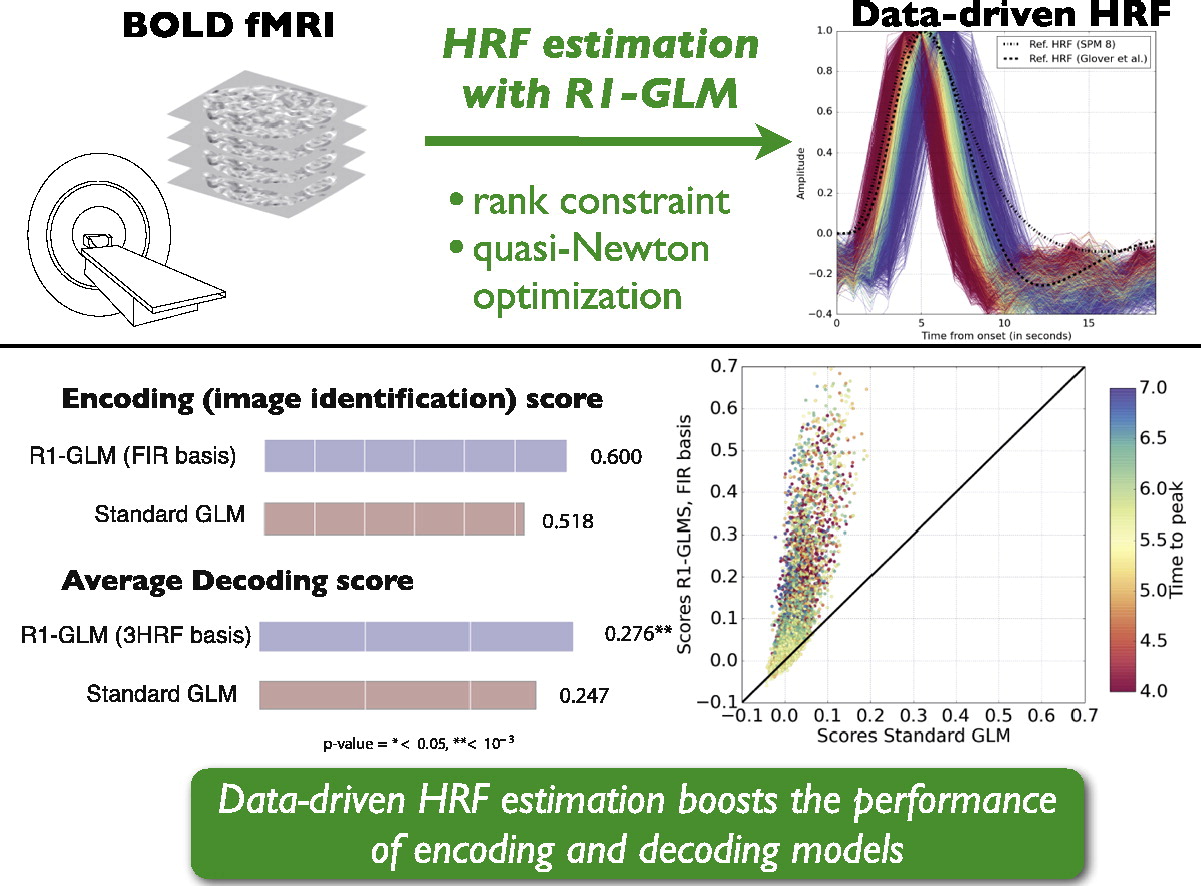
Despite the common usage of a canonical, data-independent, hemodynamic response function (HRF), it is known that the shape of the HRF varies across brain regions and subjects. This suggests that a data-driven estimation of this function could lead to more statistical power when modeling BOLD fMRI data. However, unconstrained estimation of the HRF can yield highly unstable results when the number of free parameters is large. We develop a method for the joint estimation of activation and HRF using a rank constraint causing the estimated HRF to be equal across events/conditions, yet permitting it to be different across voxels. Model estimation leads to an optimization problem that we propose to solve with an efficient quasi-Newton method exploiting fast gradient computations. This model, called GLM with Rank-1 constraint (R1-GLM), can be extended to the setting of GLM with separate designs which has been shown to improve decoding accuracy in brain activity decoding experiments. We compare 10 different HRF modeling methods in terms of encoding and decoding score in two different datasets. Our results show that the R1-GLM model significantly outperforms competing methods in both encoding and decoding settings, positioning it as an attractive method both from the points of view of accuracy and computational efficiency.
|
More details can be found in [19] .